Gene mutation guessing game/chatbot
the ideation of the gene mutation guessing game
Week 1
3/20: Original Plan: Chatbot where user inquires about a certain disease, chatbot pulls diseases and asosiated symtoms from a dataset, user answeres, chatbot provides a pertange of likleyhood of having the disease
3/21: Implementing chatbot: Utalizing gemini api key, and a dataset from symbipredict_2024, chatbot implemented in backend terminal
Week 2
3/24-3/25: Refining chatbot, implementing fuzzy searching, and completing student taught lessons/lesson homework
3/26: New Idea: Gene Guessing Game, more user input/engagment teaching about genetic conditions/causes. Creating first draft of
3/27-3/28: Finding dataset with assosiated genetic conditions:
National Institute of Health dataset , Dna sequence fetching Api
Week 3
3/31: Teach lesson to class + grade homework + implement datasets into code
4/1: Completing student taught lessons + helping with re adding header with documentation, and frontend UI (Adding Frontend to Guessing Game)
4/2: Building json file to compile data from 2 datasets, and creating API to pull both datasets together (refining search to only overlapping dieases from both sets) JSON file
4/3: Build Api fetching 2 datasets (with JSON file) to obtain the condition, gene, mutation, and sequence (making sure that the data from both files overlaps with the same gene/mutation) API
4/4: Testing get/post functions in postman, and building model file for assosiated API, saving data into SQL Lite database
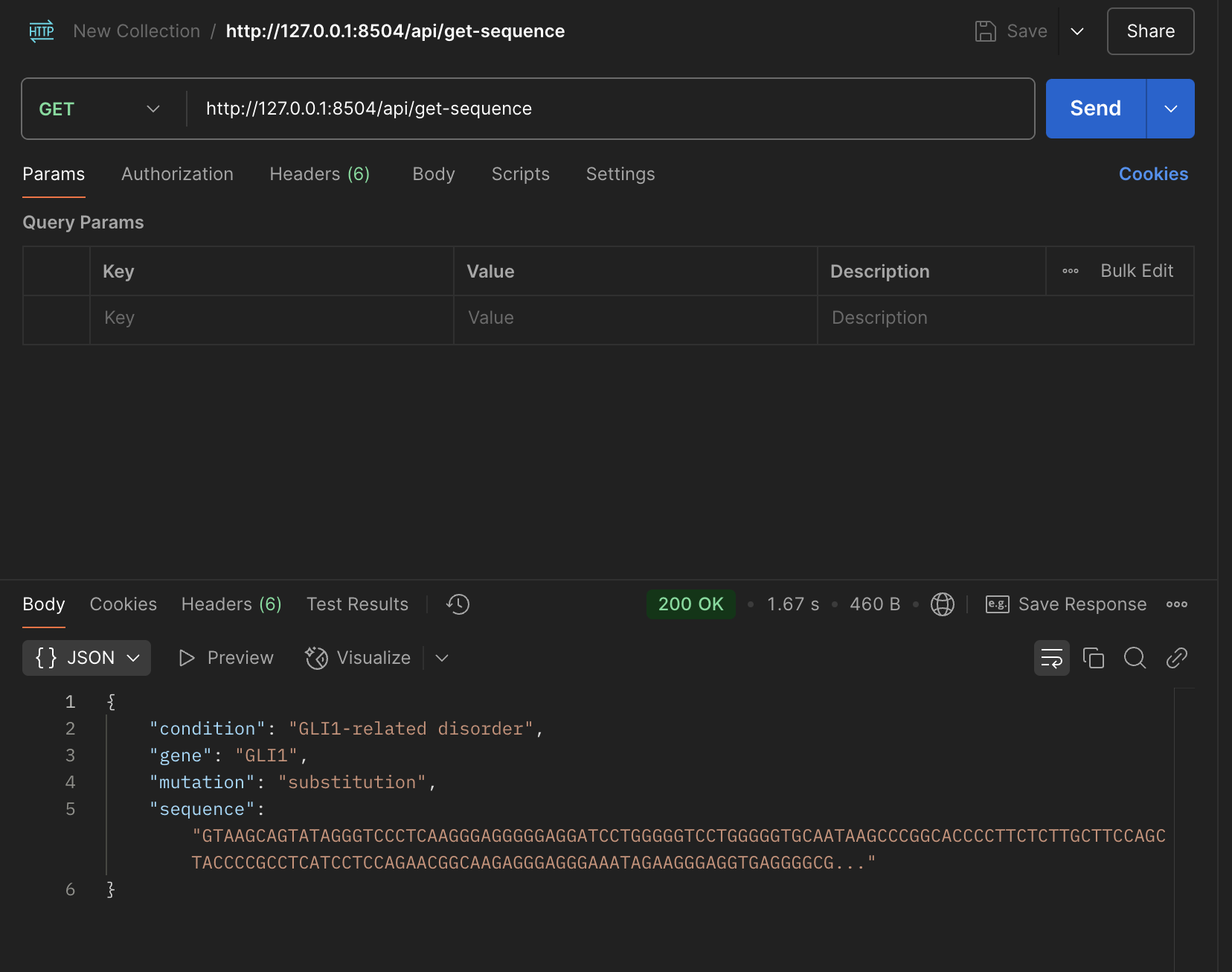 Fetching data from endpoint function in API file
Fetching data from endpoint function in API file
class GetSequence(Resource):
def get(self):
tries = 0
max_tries = 20
while tries < max_tries:
entry = random.choice(mutation_data)
gene = entry["gene"]
sequence = fetch_sequence(gene)
if sequence:
short_seq = sequence[:150] + "..." if len(sequence) > 150 else sequence
return jsonify({
"gene": gene,
"condition": entry["condition"],
"mutation": entry["mutation"],
"sequence": short_seq
})
tries += 1
print(f"[INFO] Retry {tries}: sequence not found for gene {gene}")
return jsonify({"error": "Could not find a gene with both mutation and sequence"}), 500
Explaination of function:
- fetching gene sequence randomly from mutation_data (dataset)
- If fetched correctly returns with a shortened version (150 or less charecters)
- If failed more then 20 times responds with a 500 not found error
Week 4
4/7: Student taught lessons + adding features to frontend design: color coordinates DNA sequence, button to display assosiated gene/disease, button to quiz again
4/8: Student taught lessons + fixing model/saving data correctly into SQLite database
4/9: Preparing for live review with Mr.Mort, going over agenda, features, order of presenting